Scatological science: how poo analysis could help save endangered species.

Three separate species of banded langur have been identified after DNA analysis of their faeces, paving the way for targeted conservation strategies
Across the world, conservationists, scientists and volunteers are racing to save thousands of endangered species. And for some, their efforts have not been wasted. A recent report found that conservation programmes have saved several bird and mammal species from extinction in recent decades.
But the numbers of many data deficient species and sub-species continue to dwindle – and how do you save a species if it hasn’t even been recognised?
“There are so many species for which we have no information whatsoever. If we want to conserve them, we need to know about their habitat, their food preferences, the genetic diversity and health of the population, and so on,” says Rudolf Meier, a professor at the National University of Singapore.
While DNA barcoding is increasingly becoming an invaluable tool in the identification of species, the best way to collect that DNA is still being debated.
The answer may lie in one of nature’s most basic products: poo. “In the past, we had to do two to three years of fieldwork to gather even the most basic information about a species,” says Meier. “Now, with faecal analysis, we can gather information much faster. Not only are we able to extract the host DNA, we also get to learn about its diet, gut flora and parasite load.”
It was faecal analysis that saw a recent breakthrough for Meier and his colleagues from Singapore, Indonesia and the UK in confirming three new species of banded langurs.
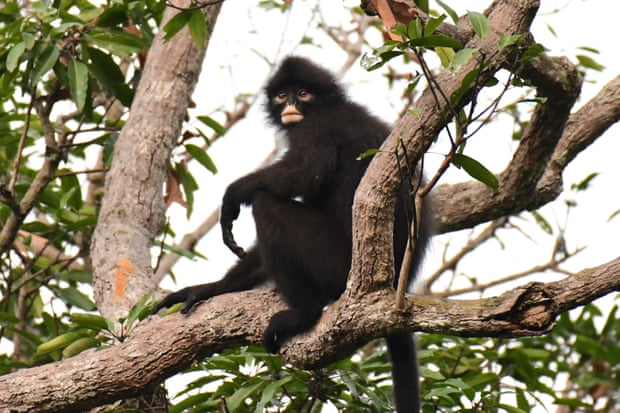
The Raffles’ banded langur and the East Sumatran banded langur were formerly considered sub-species of the langur species Presbytis femoralis. But the two south-east Asian primates have now been confirmed as species in their own right, thus qualifying them to be listed as critically endangered according to IUCN red list criteria. A third species identified by the team, the Robinson’s banded langur, is considered near threatened.
All three langurs have black bodies with only slight variations in the white markings on their faces, limbs and abdomens and so were long thought to have belonged to one fairly widespread species. They rarely descend to the forest floor and are extremely shy and elusive, making it difficult to monitor and photograph them.
The study, Faecal DNA to the Rescue, notes the difficulty in collecting data for rare species because “invasive sampling yielding fresh tissues is usually not feasible”.
“This leaves only three alternative sources of DNA,” it says. “The first is museum specimens, but the number of samples in museums tends to be small and many were collected in the 19th or early 20th century thus reflecting historical genetic diversity prior to extensive habitat loss. The second is tissue samples obtained from specimens that died of “natural causes” such as road accidents. The third source of genetic material is noninvasive samples such as hair and faeces.”
Faecal samples can be gathered during routine field surveys but it is not an easy task. It requires long hours of monitoring in sometimes harsh outdoor conditions. To ensure that an animal does not feel threatened, researchers must wait for the individual to move away after defecation before approaching the faeces.
Andie Ang, a research scientist at the Wildlife Reserves Singapore Conservation Fund and one of the paper’s co-authors adds: “It’s difficult to see the langur droppings on the soil as it is mostly brown in colour. Sometimes, when the langur defecates, the poop hits a branch or leaves [on its way down] and gets scattered all over the forest floor. Then we are looking for extremely small pieces of scat. It can get very difficult to spot.
“Sometimes, the faeces are smelly enough to be detected, other times we look out for shit flies or follow dung beetles. We are euphoric when we find a good faecal sample!”

Despite the challenges, an upside of gathering faecal samples is that it is noninvasive and does not require the animal to be captured. Unlike blood sampling or using ear clips it does not disturb or stress the animal.
For the study, DNA was extracted from the langur faecal samples using commercially available kits, and the genetic material was then sent to an external facility for sequencing. The sequenced data consisted of a minuscule amount of monkey DNA, along with DNA of the food it had consumed, parasite DNA and an overwhelmingly large amount of bacterial DNA.
Amrita Srivathsan, a researcher at the National University of Singapore and one of the paper’s co-authors says: “We isolated the langur DNA sequences using bioinformatics and complete mitochondrial genomes were reconstructed from it.”
Srivathsan says: “I had mixed feelings about the confirmation of the three banded langurs as distinct species. Their populations are small and fragmented, which is extremely worrying. But at least now, they can get the urgent attention needed for their conservation.
“The information from faecal analysis is vital in understanding the ecology of the species. It will help create targeted conservation strategies,” she adds.
In Singapore, only about 60 individuals of the Raffles’ banded langur (Presbytis femoralis) remain in one restricted forest patch, while across the Johor Strait, in southern peninsular Malaysia, numbers are also dwindling (population of 250-300 individuals), because of the impact of the rapid increase of palm oil plantations.

The East Sumatran banded langur (Presbytis percura), found in the Riau province of the Indonesian island, has lost large portions of its habitat to logging and private plantations. Small groups of the primate are now found scattered inside traditional rubber plantations, where some natural vegetation still exists. Rizaldi, a primatologist from Andalas University in West Sumatra and another of the paper’s co-authors, says: “Since we don’t have sufficient data about this species, it is not considered a protected primate in Indonesia. Our paper describing it as a species is a crucial first step towards its conservation.”
He hopes that the IUCN reassessment will bring much-needed local and international support for the protection of the species. “We cannot afford to delay,” he says.
Nadine Ruppert, head of the Primate Research and Conservation Lab at the Universiti Sains Malaysia in Penang says: “Now that there is a better understanding of the distribution boundaries and relative population sizes of the three species, more specific research and conservation action plans can be tailored for each species in their respective habitats.”
Meier, who is also one of the co-authors of the paper, feels faecal analysis is not utilised by researchers as much as it should be, something he would like to see change. “As much as we joke about ‘shitty science’ in the lab, there is an enormous amount of biological information in animal scat,” he says.
The Guardian